Research progress has helped people with Down syndrome enjoy longer lives, but scientists recognize they can still do much more to ensure that those longer lifespans are more healthful. An April 9 online symposium of the Alana Down Syndrome Center at MIT highlighted seven projects across three universities aiming to do exactly that.
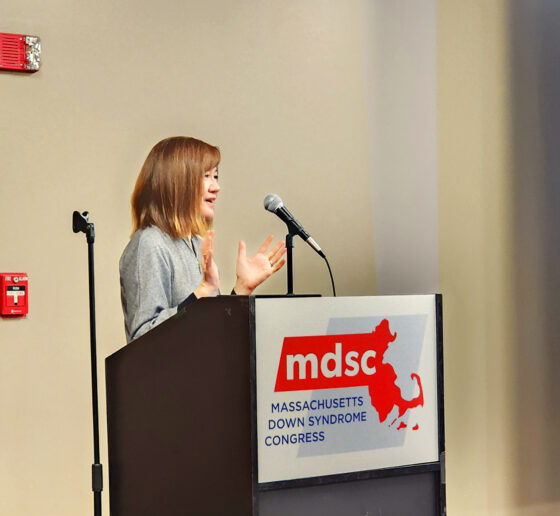
“Today, if you’re born with Down syndrome, you can expect to live 60 or more years. There are people with Down syndrome that have lived well into their 80s,” said Rosalind Firenze, scientific director of the Alana Center based in The Picower Institute for Learning and Memory at MIT. “But unfortunately, when we think about the health of people with Down syndrome, research into medicine and health has been historically underfunded, and often people with Down syndrome are excluded from research.
“So there are many unknown things about Down syndrome biology and neuroscience, but it’s really important to search for answers, because what we do know is that our loved ones with Down syndrome have a heightened risk for a number of conditions,” Firenze added.
Mindful of these disparities, Picower Professor Li-Huei Tsai and colleagues founded the Alana Down Syndrome Center (ADSC) at MIT in 2019 with the generous support of the Alana Foundation. Its mission, Firenze said, is to “build research, training and technology for a world of greater inclusion, possibility and accessibility for people with Down syndrome.”
In the symposium “Accelerating Research for Down Syndrome,” three ADSC supported fellows presented their projects on brain and heart development alongside four researchers from Rutgers University and the University of São Paolo (USP) in Brazil, who discussed their studies of cognition and brain health over the lifespan.
Brains young to old
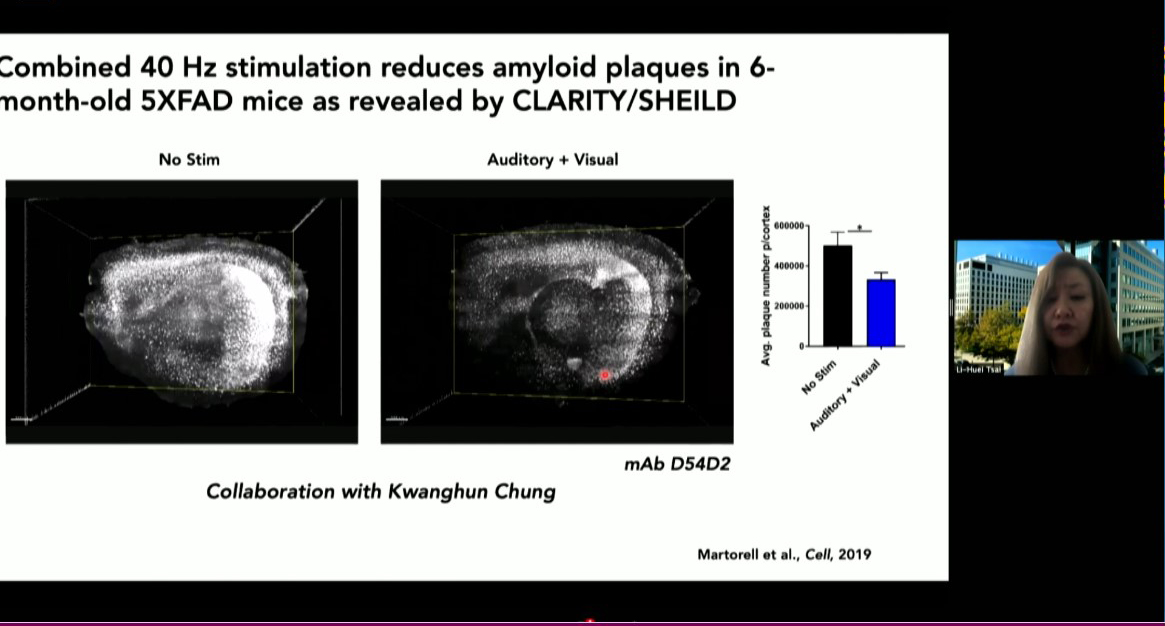
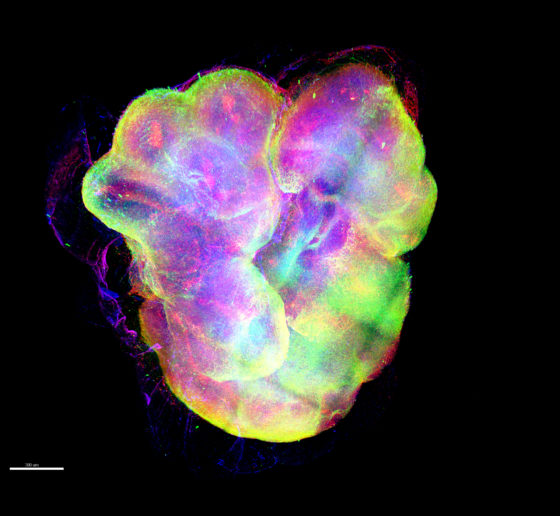
In 2024, Tsai’s Picower Institute lab showed that in mice exposed to light flickering and sound clicking at the 40Hz frequency of gamma brain waves, specific neurons increased their output of the peptide VIP. Other studies indicate the peptide has important roles in fetal brain development. In her symposium talk, Alana Fellow and postdoc Roberta Fonseca described how members of the lab have begun testing whether exposure to the sensory stimulation among pregnant mice increases VIP in the womb and benefits brain development of their fetuses in a model of Down syndrome. They are also testing whether applying VIP to developing human stem cell cultures with trisomy 21 enhances their survival and metabolic health. The lab is still working on the experiments, including memory tests for Down syndrome mice born from mothers that were exposed to the 40Hz stimulation, compared to unstimulated controls.
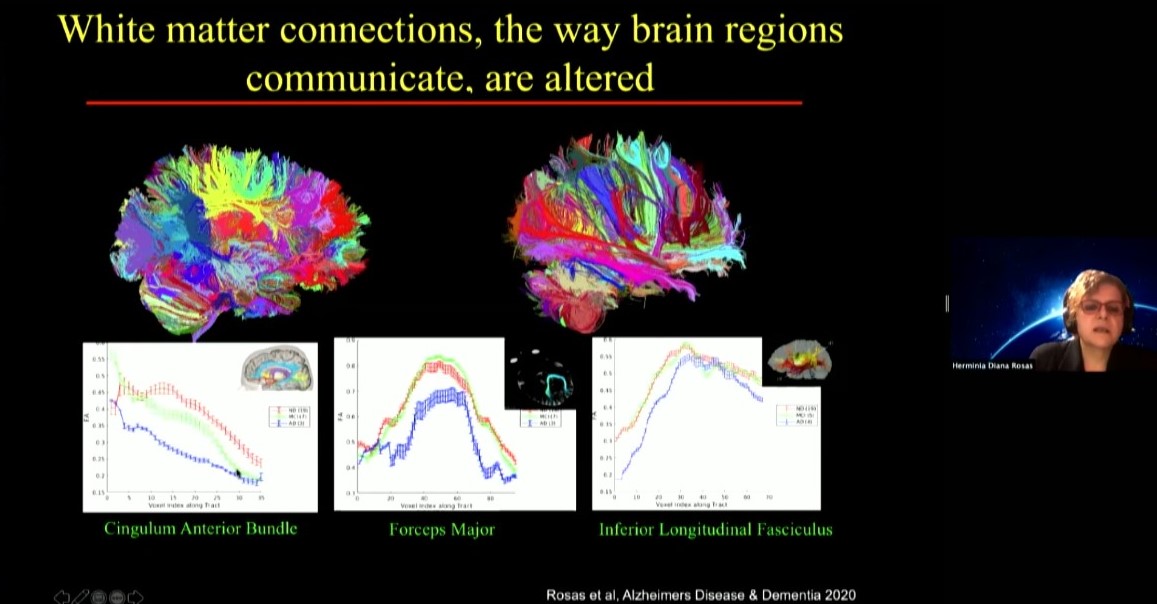
USP Professor Daniele De Paula Faria presented her work on brain imaging of both people with Down syndrome and mouse models at ages old and young. She’s using positron emission tomography (PET) scanning to track key molecular indicators of metabolism and inflammation. In a study she published last year in Alzheimer’s & Dementia, for instance, she and colleagues showed that people with Down syndrome show significantly greater neuroinflammation compared with neurotypical people as young as age 20 and that the increased inflammation correlates strongly with amyloid plaque, which are well-known biomarkers of Alzheimer’s disease. She has seen similar indications in mice and has begun testing potential interventions.
Also looking across the lifespan for predictors of Alzheimer’s development, Rutgers Assistant Professor Luciana Fonseca described her work studying how cognitive variability at a single point within individuals with Down syndrome can provide important information about AD risk and resilience. Rather than capturing sparse snapshots of cognition, as many clinicians do, she is leading a study in which her team assesses cognitive performance across tasks and tests to enable measurements of variability. Her hypothesis is that the greater the degree of variability, the higher the risk of developing dementia. In the study she’s also collecting other potential biomarkers associated with Alzheimer’s onset.
Rutgers graduate student Chuhanwen Sun, meanwhile, presented his research aimed at understanding at the cellular level what makes the brain vulnerable to Alzheimer’s disease in Down syndrome. Using postmortem brain samples from Down syndrome individuals who did or did not develop Alzheimer’s, he has engaged in a deep analysis of gene expression and epigenetic differences in individual cell types. The work is still ongoing but he said the research is pinpointing specific cells in a specific brain region as being especially vulnerable.
Since 2019, USP Professor Orestes Forlenza, a geriatric psychiatrist, has been enrolling hundreds of adults with Down syndrome in a study aimed at better understanding how various blood-based biomarkers track with changes in cognition over the lifespan. Among these are markers of cellular oxidative stress, measures of cell death or “apoptosis,” and a form of molecular signaling called the wnt pathway. He’s also been able to rule out other suspected biomarkers that don’t correlate with cognitive impairment in Down syndrome volunteers.
The developing heart
Two postdocs supported by the ADSC and working in the lab of MIT Biology Professor and ADSC investigator Laurie Boyer talked about their research on the heart. Their goal is to identify factors at play in the earliest stages of development to better understand, and potentially intervene to prevent, congenital heart defects that are much more common in people with Down syndrome than in the general population.
Alana Fellow Mehdi Hamouda investigates how physical forces on developing heart cells may differ in trisomy 21 vs. in cells that have the typical two chromosome copies. The forces on cells can impact the nucleus, affecting how stem cells ultimately become more specific cells in the body, including the cardiomyocytes that build the heart. His hypothesis is that trisomy 21 causes faulty mechanical cues that change how the nucleus feels and responds to these forces to impact the earliest stages of heart development.
Meanwhile, NRSA postdoc Leah Borden presented her work integrating both mouse hearts from Down Syndrome models and trisomy 21 cells from individuals with DS. Her research investigates how developing heart cells engage with their surrounding environment, the “extracellular matrix” (ECM), to regulate critical tissue properties that guide differentiation. Leah is uncovering how Trisomy 21-associated alterations in ECM reshapes the mechanical and signaling landscape during heart development.
In all, the symposium provided seven examples of how ADSC researchers and their colleagues are racing to better understand factors that affect the lifelong health of people with Down syndrome.