Author: Emily Niederst
Technology to Improve Ability program funds sleep apnea device

In collaboration with the Alana Down Syndrome Center at MIT, the Deshpande Center for Technological Innovation offers a grant program: Technology to Improve Ability (TTIA).
The mission of the TTIA program is to commercialize technology that will improve the quality of life for people with Down syndrome and other disabilities. The Deshpande Center assists MIT faculty and students in commercializing their technologies, taking promising ideas and innovations and turning them into products and services. While the focus is to help people with Down syndrome, the innovations are also likely to aid a much broader population.
The TTIA program continues to fund a project to create a better means of treating obstructive sleep apnea for people with Down syndrome. The project team was funded for its second year through the TTIA program.
Obstructive sleep apnea (OSA) is a common, chronic health condition in which repetitive collapse of the throat muscles results in episodic obstruction of breathing called apnea. The most common location for obstruction is where the tongue meets the soft palate at the back of the throat. OSA results in poor sleep quality and is associated with serious health consequences. It is common in the Down syndrome (DS) population, affecting 50% of children and almost all those in adulthood.
OSA Project Team leadership

The device combines a customized small oral prosthesis with low suction to hold the tongue to the top of the mouth, clearing the airway. Standard treatment for sleep apnea is a continuous positive airway pressure (CPAP) machine, which requires users to wear a mask at night while a pump pushes air into their nose and/or mouth. The masks are difficult to fit, particularly for people with non-standard face shapes, are not very comfortable, and the pump is loud. Half of CPAP users discontinue use by a year, and users with DS have significantly lower rates of compliance. The need for an alternative treatment is great.
In the past year, the team refined designs for its mouth prosthesis and began testing the comfort and wearability of the device in a preliminary trial with MIT volunteers. The results will set the stage for a larger clinical trial to be conducted early next year among patients with sleep apnea at Boston Veterans Affairs Healthcare.
New tools for dynamic analysis and molecular mapping of the brain
Dr. Ed Boyden’s lab has been building tools for the dynamic analysis and molecular mapping of the brain for all Alana Center groups to better understand the differences present in Down syndrome.
Imaging neurons as they fire
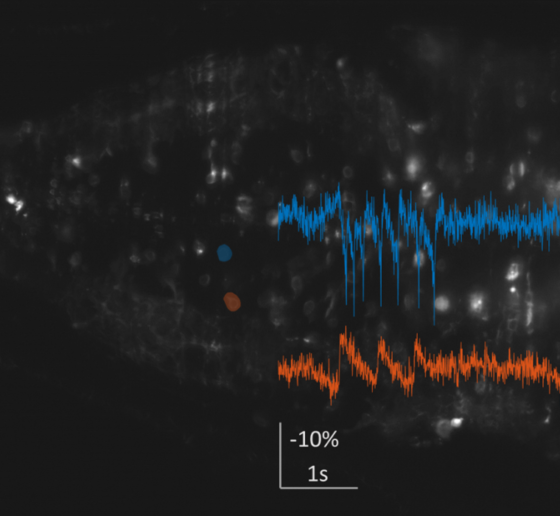
In a first line of work, the Boyden Lab has been developing imaging tools to characterize brain activity in Down syndrome model mice at the high speeds necessary to understand the mechanisms underlying the Tsai lab’s 40 Hz sensory stimulation protocol, GENUS. To do so, the lab has been identifying and developing new voltage indicators and creating new microscope architectures. The lab identified an excellent voltage indicator for in-animal neuron voltage imaging. To make high-speed, 3D imaging possible in the brains of freely moving mice, they are designing a miniaturized light-field endoscope that will allow simultaneous recording of neural activity in a 3-D volume at exceptionally high imaging rates. They have fabricated customized components, are working to finalize its construction, and are planning to apply it to studying Down syndrome mouse models in collaboration with the Tsai lab.
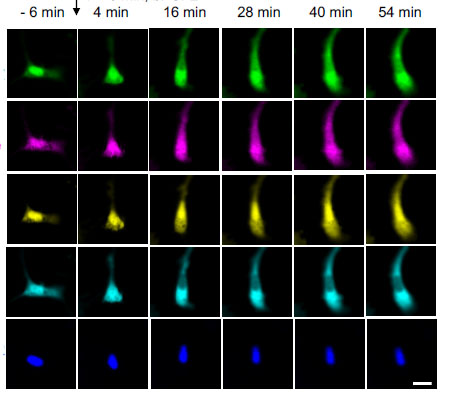
As part of this work, the Boyden lab recently identified sets of fluorescent probes that could be utilized simultaneously to do massively parallel imaging of many signals at once in a living cell, which is key to understanding how cells compute.
Expanding brains to see more clearly
Down syndrome involves changes in the organization of neural connections called synapses, which are too small scale to be imaged. In a second line of work, the Boyden Lab is refining its technology of expansion microscopy (ExM), which physically enlarges objects so they can be imaged with nanoscale precision on ordinary lab equipment.
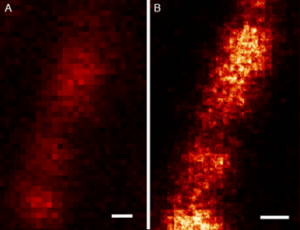
They are developing a more powerful chemical method that locks proteins in place while enabling them to be cleanly separated from each other. The team is also creating a freezing protocol that uses cold temperatures to lock molecules in place while the aforementioned process is fixing them. The result is better preservation of delicate nanostructures. As part of this work, the team was able to publish their procedure on expanding proteins away from each other to greater extents than previously possible (see article on MIT news).
“Big Data” to probe intersections of neurological disease
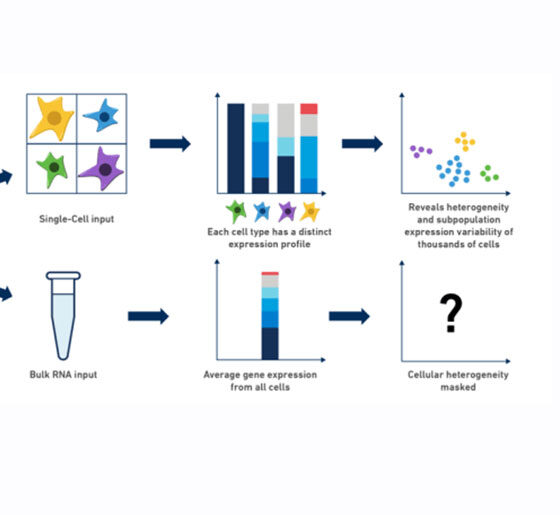
Down syndrome changes how an individual’s body and brain develop, which can cause mental and physical challenges later in life. However, outcomes vary greatly across individuals and their cognition, behavior, and social interactions. The molecular underpinnings of such variability remain unknown and could make a great difference for therapeutic interventions that can ameliorate outcomes for many individuals with Down syndrome. Understanding the molecular basis of such differences requires systematic mapping of the molecular pathways, biological processes, regulators, individual cells, and brain regions in the context of multiple individuals with Down syndrome with diverse outcomes.
Due to an extra copy of the Alzheimer’s-related gene APP on chromosome 21, cognitive decline and amyloid plaque formation characteristic of Alzheimer’s disease are also greatly increased in prevalence across individuals with Down syndrome, providing an important and well-suited setting for studying phenotypic, molecular, cellular, and biological variability.
To address the heterogeneity of biological processes, cell types, brain regions, and individual cells across individuals with Down syndrome, Alana Center member Manolis Kellis has profiled 135,597 cells from 24 donated human postmortem samples with Down syndrome, some with and some without Alzheimer’s Disease. The team looked across two brain areas at the types and health of cells found in those regions, as well as the profile of gene expression found in those cells.
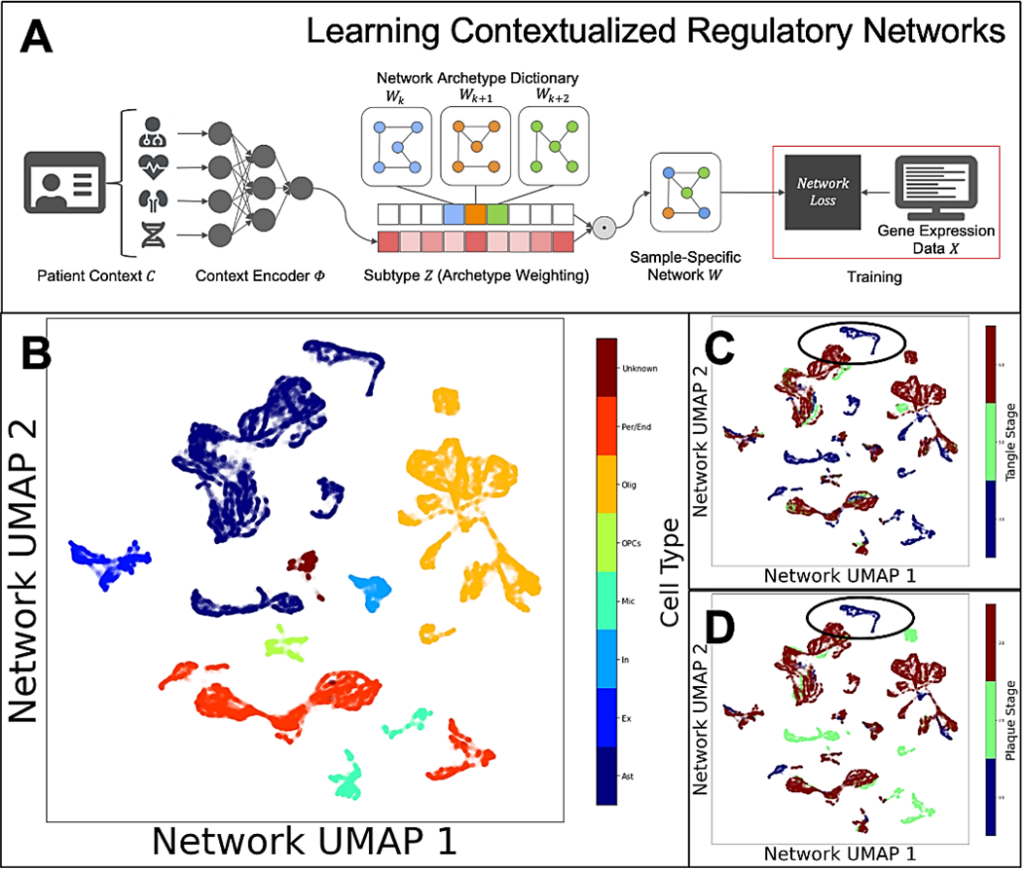
To do this, the team generates an ‘single-cell atlas,’ and labels and clusters cells into cell types, and then looks for genes that are regulated differently between the cell types. The lab also uses sophisticated computational methods to examine how gene expression is regulated in different cell types and states, with the goal of understanding the regulatory networks that control these. Gene regulatory networks are usually highly structured and regulated by a limited set of active regulators, which make them attractive candidates to approach as treatment targets that may cover many altered cellular processes.
Exciting preliminary discoveries
The lab has made some preliminary discoveries. They found that the presence of Alzheimer’s disease is associated with significant decreases in astrocytes and endothelial cells in a frontal cortical brain region. They observed down-regulation of DYRK1A, a gene previously linked to intelligence found on chromosome 21, in a subset of excitatory neurons in the amygdala.
The team is also studying altered gene regulatory networks in astrocytes. Alana fellow Ben Lengerich developed computational methods to address inter-individual and inter-cell diversity by estimating gene regulatory networks influenced by patient-specific, tissue-specific, disease-specific, and cell-specific contexts and found that astrocyte networks cluster according to the onset of tangle and plaque formation.
Lastly, Alana fellow Bo Zhao and former fellow Michael Gutbrod developed high-throughput methods to alter gene regulators in iPSC-derived neurons and microfluidics-based spatial transcriptomics.
Tsai lab expands Down syndrome research

ADSC Director Li-Huei Tsai‘s lab has significantly expanded their Down syndrome (DS) work. New research is looking at brain oscillations and sleep in DS model mice versus neurotypical mice. Continued research on using 40Hz light and sound (GENUS) therapy for several weeks in DS mice has revealed improvements including producing new neurons in the adult brain and reducing brain immune cells. The lab is investigating performance on memory tasks and has also launched a new project testing the benefits of GENUS during pregnancy on development of fetuses with trisomy 21. Finally, the Tsai lab has begun a Phase 1/2 human clinical study with GENUS light and sound devices in individuals with DS. The treatment appears to be safe and well-tolerated with no adverse effects so far.
40Hz light and sound in sleep and neurogenesis
By the age of 40, most individuals with DS present a substantial accumulation of amyloid beta (Aβ) plaques and neurofibrillary tangles (NFT), toxic protein deposits considered to be hallmarks of Alzheimer’s disease. Pioneering studies performed by the Tsai laboratory have shown that sensory stimulation (i.e., light and sound) at the frequency of a key brain rhythm (gamma: 40Hz), a technique called GENUS, enhances gamma rhythms in the brain and can have neuroprotective effects in the context of Alzheimer’s disease. Recent work from the Tsai lab suggests this to be true in mouse models of DS as well. Recent work suggests a key mechanism through which GENUS exerts a neuroprotective effect is by promoting the clearance of waste products through the glymphatic system. The strongest known regulator of glymphatic clearance of waste is sleep. Therefore, a new research project aims to investigate whether GENUS can be performed during sleep and to assess how the potential neuroprotective effects of this treatment compare to the effects of using GENUS during wakefulness. The results of this project could provide a pathway to enhance the beneficial effects of GENUS in the context of Alzheimer’s disease and DS. Moreover, the ability to deliver GENUS during sleep in future clinical trials or interventions would lead to a striking increase in compliance with the treatment and the daily amount of stimulation, particularly for individuals with DS. Taken together, the administration of GENUS during sleep promises a striking increase in treatment efficacy.
 To address this aim, Alana Fellow Cristina Blanco-Duque has built a setup that supports continuous (24h/7 days a week) electrophysiology recordings in mice and is equipped to perform GENUS. She also established multi-site in vivo electrophysiology recordings in a mouse model for DS. This setup has already produced several rich datasets with exciting results, including that the DS model mice show fragmented and lighter sleep than their wild-type littermates, and may have hyper-synchronized brain rhythms.
To address this aim, Alana Fellow Cristina Blanco-Duque has built a setup that supports continuous (24h/7 days a week) electrophysiology recordings in mice and is equipped to perform GENUS. She also established multi-site in vivo electrophysiology recordings in a mouse model for DS. This setup has already produced several rich datasets with exciting results, including that the DS model mice show fragmented and lighter sleep than their wild-type littermates, and may have hyper-synchronized brain rhythms.
Blanco-Duque and Tsai lab colleagues have also developed a stimulation system that allows the lab to deliver GENUS selectively during sleep or wake. This setup will be used to test whether delivery of the GENUS light and sound treatment is possible during sleep, and how this affects sleep/wake patterns. This system can also be used to study the effects of GENUS on the clearance of toxic waste products during sleep.
The lab has also found in early results that after three weeks of 40 Hz light and sound (GENUS) stimulation in the mouse model of DS there is a striking benefit of the production of new neurons in the adult brain. This process has been shown previously to be significantly impaired in mouse models of DS, which may contribute to some of the cognitive dysfunction that has been observed. The lab is also investigating the effects of extended GENUS treatment on several behavioral tasks, including spatial short-term memory and anxiety behaviors, as well as markers of inflammation and immune activity in the brain.
Based on preliminary data using single-nucleus RNAseq – a high-resolution method of quantifying gene expression in individual brain cells – the lab hypothesizes that GENUS may alter the expression of several critical DS-related genes that influence the balance between the production of new astrocytes and neurons.
GENUS human clinical studies

Work has started within the Tsai lab to translate the findings in mice into benefits for human subjects with DS. Clinical work began by investigating the safety, compliance, and entrainment of the 40 Hz stimulus. In a recent publication, the Tsai lab found that repeating auditory tones can also be used to entrain gamma oscillations, and in combination with visual stimulation, GENUS can impact brain regions outside of the primary sensory cortex and extend into the hippocampal memory. With combined visual and auditory stimulation at 40Hz, the entrainment of gamma oscillations can be seen across the auditory cortex, visual cortex, hippocampus, and medial prefrontal cortex. Based on these studies, the team developed a non-invasive medical device that will possibly be effective in preventing the progression of AD and other pathology in individuals with DS. Our objective is to determine whether non-invasive sensory stimulation can be used to modulate gamma power and synchronization in individuals with DS as a potential therapeutic to prevent AD and other cognitive troubles in this population.
In a first Phase 1/2 study, the team will treat 30 individuals with DS and age-matched, cognitively typical controls with the GENUS light and sound device while using electroencephalography (EEG) to evaluate induced entrainment and effects on their brain circuitry. Participants are blindly randomized to receive 1-hour of either sham or active 40Hz light and sound stimulation. The team performed cognitive testing before and after the stimulation session. Data so far suggests that our prototype GENUS light and sound device is safe and tolerable in participants with Down syndrome with no significant adverse effects and no evidence of seizure-like activity on EEG during treatment. The lab is continuing to recruit study subjects , with approximately 18 adults with DS already participating in the study.
Prenatal GENUS
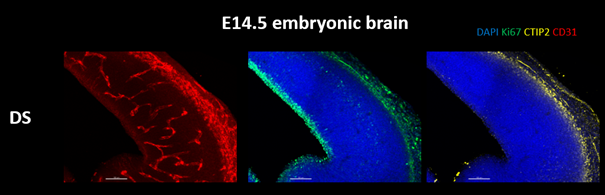
New work in the Tsai lab is finding that the effects of GENUS extend beyond the brain. GENUS stimulation in aged mice changes the number of intestinal immune macrophages and the microbiome composition of the gut. It has been previously shown that immune changes in pregnancy can have large effects on the fetus. Therefore, we started a new project in the lab to understand if GENUS treatment during pregnancy can improve deficiencies found in the brain development of Ts65Dn embryos. The lab is investigating changes in the reduction in brain size and cortical plate thickness observed in Ts65Dn embryos after maternal GENUS treatment. The lab will analyze the maternal plasma, placenta, and brain to understand the possible mechanisms by which GENUS ameliorates the embryonic brain. They also aim to understand if prenatal GENUS treatment can benefit the offspring postnatally in terms of their brain development and function.
Symposium examines intersecting biology of neurodegeneration, Down syndrome
Talk Videos
Watch videos of several of the symposium talks on YouTube.
Neuroscientists still have a tremendous amount to learn about the causes and courses of neurodegenerative diseases and Down syndrome, but as speakers at the Oct. 5-6 MIT symposium “Glial and Neuronal Biology of the Aging Brain” pointed out, often when they make a new discovery in the context of one such condition, it teaches them something relevant to others.
“Our belief is that the study of the aging brain can learn a great deal from the study of Down syndrome and vice versa,” said Picower Professor Li-Huei Tsai who directs the two MIT entities that jointly hosted the conference: The Aging Brain Initiative and the Alana Down Syndrome Center. “It would be a wonderful outcome of this symposium if we can play even a small role in bringing these two communities of scientists, physicians, and engineers, and even caregivers closer together.”
The event indeed marshaled a multitude of online attendees. Over the course of the two-day program more than 400 people tuned in from 27 countries. They heard scientists from places as far-ranging as Hong Kong and Germany share their latest research and discuss the many intersections they see among Alzheimer’s and other dementias, Parkinson’s disease, Huntington’s disease and Down syndrome.
For example, Tracy Young Pearse, associate professor of neurology at Harvard Medical School and Brigham and Women’s Hospital, discussed her lab’s new finding that the three copies of the genes APP and DYRK1A found in Down syndrome neurons (because they have three copies of chromosome 21), increase phosphorylated tau (a pathological hallmark of Alzheimer’s) and promote excessive transport and release of neurotransmitters across connections with other neurons, a potential source of circuit dysfunction.
Vessels of concern
Though neural circuits remain at the heart of brain function, three speakers instead focused their talks on the brain’s circulatory system. MIT Associate Professor Myriam Heiman noted that the breakdown of the blood-brain barrier, which strictly filters what the body and brain exchange, are suspected of being key contributor to many neurodegenerative diseases. In presenting her lab’s new research that produced a novel “atlas” of cell types in the brain’s blood vessels, she showed clear evidence that vascular integrity is weakened in Huntington’s disease and that the degradation is associated with a problematic innate immune response.
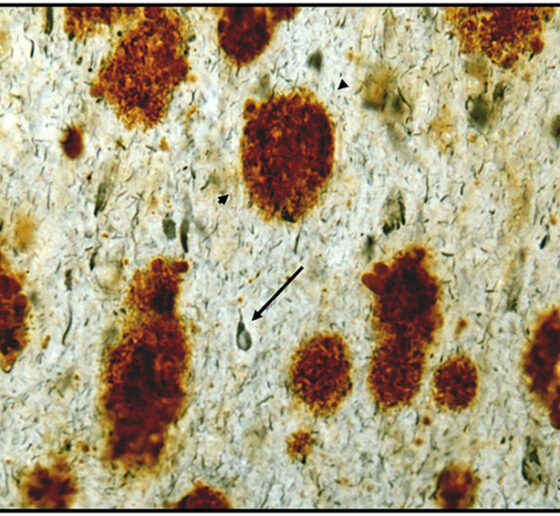
Elizabeth Head, Professor of pathology and laboratory medicine at the University of California at Irvine, related dysfunction of brain vasculature to the connection between Down syndrome and Alzheimer’s. Though people with Down syndrome are relatively protected against cardiovascular problems such as high blood pressure or atheroma, an excess of amyloid protein in their brain blood vessels leads to cerebral amyloid angiopathy, a condition closely associated with Alzheimer’s. Head’s lab has shown that people with Down syndrome and CAA exhibit microbleeds along their brain blood vessels.
Head collaborates with Adam Brickman, professor of neuropsychology at Columbia University. He presented recent studies showing that magnetic resonance imaging of “white matter hyperintensities” and other vascular problems can be a biomarker of Alzheimer’s pathology in people with Down syndrome. The hyperintensities, which the team showed to be especially prevalent in posterior lobes of the brain, are believed to be the result of brain vasculature problems and correlated with other problems such as microbleeds.
Cells not immune from scrutiny
Several other speakers focused on the brain’s immune cells, called microglia, which have a very complex role in neurodegenerative diseases including Alzheimer’s.
Microglia, for instance, take on many different states in Alzheimer’s ranging from beneficial to harmful. In her talk, Harvard Medical School & Boston Children’s Hospital neurology Associate Professor Beth Stevens described methods her lab has developed for culturing microglia from stem cells and then coaxing them into these many states by tailoring either their genetic background, their environmental context, or both.
Li Gan, professor of neuroscience at Weill Cornell Medicine, discussed particular instances in which molecularly manipulating microglial state can sustain the brain’s resilience to Alzheimer’s pathology. In a study published earlier this year her lab found that by reducing expression of the gene transcription factor NFkappaB in microglia, the lab could reduce spreading of the problematic protein tau. She also shared even newer results showing that intervening in a specific runaway immune pathway in microglia by knocking down a key molecule, her lab has shown benefits in learning and memory in mice. The method appears to do so by increasing activity of a resilience-promoting transcription factor called MEF2 that Tsai’s lab has also independently identified as beneficial.
Hong Kong University of Science and Technology Professor Nancy Yuk-Yu Ip detailed another molecular method of helping microglia combat Alzheimer’s. Her lab has found that in the disease a soluble form of the molecule ST2 intercepts the immune molecule interleukin 33 (IL-33), which would normally prompt a transition of microglia into a beneficial state. The lab has shown that injecting IL-33 improves Alzheimer’s pathology in mice and has found a genetic variant in people that conveys protection against this problem.
In his talk, Michael Heneka, director of the Luxembourg Centre for Systems Biomedicine, showed how microglia literally throw neurons a line to help them fight back against toxic proteins. His lab found that microglia extend “tunneling nanotubes” to neurons beset with tau (a toxic aggregate in some dementias) or alpha-synuclein (a toxic aggregate most prevalent in Parkinson’s disease) to remove the proteins and to supply neurons with fresh mitochondria to rescue them from oxidative stress.
A system with many parts
Neurons, vascular cells, and microglia were not the only cells with time in the spotlight. Shane Liddelow, assistant professor of neuroscience and physiology at New York University focused on astrocytes, an abundant cell type in the brain with key roles in supporting neural function and linking neurons to blood vessels. He shared new research indicating that subtypes of astrocytes have inflammatory responses in disease and in the case of Alzheimer’s, associate with pathology in particular parts of the brain. Further research can help determine what those subtypes may matter to the progression of the disease.
Astrocytes, neurons and microglia were all featured in the remarks of Gilbert Di Paolo, executive director of discovery biology at Denali Therapeutics. He discussed the company’s potential therapy for a subset of cases of frontotemporal dementia. In those cases, mutations reduce levels of progranulin, which undermines the function of cells’ lysosomes. By restoring levels of progranulin in cells the company is restoring lysosomal function and therefore indicators of cell health.
Complementing the talks’ exposition of the variety of cell types and molecular mechanisms at issue across neurodegenerative diseases and Down syndrome were the posters of MIT postdocs and graduate students that followed the talks. A dozen presenters from seven labs affiliated with the Aging Brain Initiative, the Alana Center, or both highlighted whole systems approaches to understanding and treating disease. Members of Tsai’s lab, for instance, discussed the therapeutic possibilities for Down syndrome of stimulating the brain with light and sound at the key frequency of 40Hz. Members of the labs of Professors Ed Boyden and Alan Jasanoff presented new advances in brain imaging. Members of Professor Manolis Kellis’s lab showed how sophisticated computational approaches can help demystify the genetic complexities of Down syndrome. A poster representing the lab of Professor Ernest Fraenkel highlighted molecular networks related to neurodegeneration. And members of the labs of Professors Ann Graybiel and Matthew Wilson highlighted neural mechanisms fundamental to behavior and memory.
The symposium offered all these scientists, and their hundreds of audience members, the chance to virtually gather and learn from each other at a crossroads of intersecting disease biology.
In memoriam of Professor Tom Hehir

The Alana Down Syndrome Center honors the memory of Professor Thomas Hehir, a long-time advocate of students with disabilities. He died on Wednesday, June 22, after a battle with ALS. Hehir had a long and storied background working in education, most recently as a faculty member at the Harvard Graduate School of Education. He was an ally in the defense of human rights, particularly children and people with disabilities, and was a staunch supporter of integration and inclusion of all students together in schools. His career spanned policy, writing, teaching, public service, and was an contributor to the Alana Institutes efforts to advocate for the rights of students with disabilities.
“Thomas Hehir was one of the easiest people to love I’ve ever met. He worked tirelessly so that schools are spaces for all children and adolescents, and he inspired Alana in several initiatives to strengthen an education that values differences”, comments Ana Lucia Villela, president of the Alana Institute and Foundation.
In 2016, Hehir coordinated an Alana Institute survey on the benefits of inclusive education for students with and without disabilities. He was also featured in the film “Forget Me Not,” produced by Maria Farinha Films and Route 6. Hehir’s work in this area helped inspire the MIT ADSC statement of purpose and goals.
He regarded the fact that many students with disabilities are still studying in separate environments as a “great failure.” His legacy reinforces the commitment to a profound transformation in schools so that they can respond positively to the diversity of students. May he be an example for those striving for a society where no one is left behind.
Upcoming Alana Down Syndrome/Aging Brain Initiative Symposium
The topic of this symposium is Glial and Neuronal Biology of the Aging Brain
REGISTER HERE:
-
Adam M. Brickman, Columbia University
-
Gilbert Di Paolo, Denali Therapeutics
-
Li Gan, Weill Cornell Medical College
-
Elizabeth Head, University of California, Irvine
-
Myriam Heiman, Picower Institute, MIT
-
Michael Heneka, Luxembourg Centre for Systems Biomedicine (LCSB)
-
Nancy Yuk-Yu Ip, The Hong Kong University of Science and Technology
-
Shane Liddelow, NYU
-
Beth Stevens, Boston Children’s Hospital, Harvard Medical School
-
Tracy Young-Pearse, Brigham and Women’s Hospital, Harvard Medical School